Emma C. Thompson, Laura E. Rosen, James G. Shepherd, Robert Spreafico, Ana da Silva Filipe, Jason A. Wojcechowskyj, Chris Davis, Luca Piccoli, David J. Pascall, Josh Dillen, Spyros Lytras, Nadine Czudnochowski, Rajiv Shah, Marcel Meury, Natasha Jesudason, Anna De Marco, Kathy Li, Jessia Bassi, Aine O’Toole, Dora Pinto, Rachel M. Colqohoun, Katja Culap, Ben Jackson, Fabrizia Zatta, Andrew Rambaut, Stefano Jaconi, Vattipali B. Sreenu, Jay Nix, Ivy Zhang, Ruth F. Jarrett, William G. Glass, Martina Beltramello, Kyriaki Nomikou, Matteo Pizzuto, Lily Tong, Elisabetta Cameroni, Tristan I. Croll, Natasha Johnson, Julia Di Iulio, Arthur Wickenhagen, Alessandro Ceschi, Aoife M. Harbison, Daniel Mair, Paolo Ferrari, Katherine Smollett, Federica Sallusto, Stephen Carmichael, Christian Garzoni, Jenna Nichols, Massimo Galli, Joseph Hughes, Agostino Riva, Antonia Ho, Marco Schiuma, Malcolm G. Semple, Peter J. M. Openshaw, Elisa Fadda, J. Kenneth Baillie, John D. Chodera, The ISARIC4C Investigators, the COVID-19 Genomics UK (COG-UK) consortium, Suzannah J. Rihn, Samantha J. Lycett, Herbert W. Virgin, Amalio Telenti, Davide Corti, David L. Robertson, and Gyorgy Snell.
Cell 184:1171, 2022. [DOI] [PDF] [bioRxiv] [Supplementary Info] [Folding@home data]
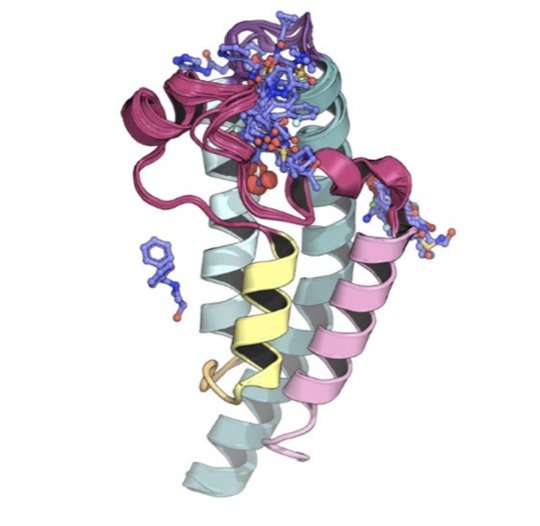
New mutations that enhance the affinity of SARS-CoV-2 spike protein for human ACE2—and potentially pose threats to antibody-based therapeutics and vaccines for COVID-19—are already emerging in the wild. We characterize and describe sentinel mutations of SARS-CoV-2 in the wild that herald challenges for combatting COVID-19, and use simulations of the RBD-ACE2 interface on Folding@home to biophysically characterize why these mutations can lead to enhanced affinity.