Computational datasets
Explicit solvent Folding@home trajectories for Aurora A kinase (AurA) from our Nature Chemical Biology paper
Dataset from:
Soreen Cyphers, Emily F Ruff, Julie M Behr, John D Chodera, and Nicholas M Levinson.
A water-mediated allosteric network governs activation of Aurora kinase A
Nature Chemical Biology, in press. [DOI] [GitHub]
We have made all the explicit-solvent Folding@home simulation data and analysis scripts used in this paper available for download:
http://github.com/choderalab/AurA-materials
The trajectory data itself is too large to share via GitHub, so we make it available via the Open Science Framework.
A microsecond trajectory of the human tyrosine kinase ABL1 kinase domain generated on Foldign@home using the OpenMM 6.3 core (core 21).
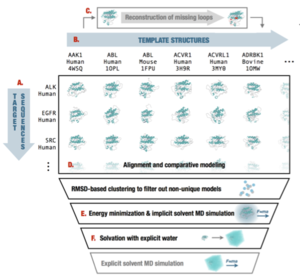
Ensembler-generated Tyrosine kinase models
A collection of structures of all human tyrosine kinase catalytic domains modeled onto all kinase domain structures from any organism from the RCSB using our Ensembler high-throughput modeling and simulation setup tool.
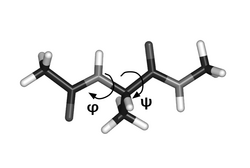
Solvated alanine dipeptide - thermodynamics and kinetics
A collection of datasets of the alanine dipeptide (more specifically, terminally-blocked alanine peptide) in explicit solvent. These datasets cover both thermodynamic simulations (parallel tempering) and kinetic simulations (Hamiltonian dynamics trajectories) useful for testing algorithms analyzing the thermodynamics and kinetics of biomolecular systems. This model system has already been used in several research papers.
Experimental datasets
Laboratory automation system data feeds
Temperature/humidity sensor feeds for Cytomat Hotel and Tecan EVO deck.
Human kinome bacterial expression data
Constructs of human kinase domains known to express well in E. coli, using the PDB as a source of potential expression constructs, confirmed by automated cloning and expression.
Manuscript prior to publication: [bioRxiv]
Interactive data browser: [github.io]
Plasmids available via AddGene
Cyclohexane / water distribution coefficients for the SAMPL5 challenge
For the purpose of the SAMPL5 distribution coefficient challenge, 53 small-molecule cyclohexane/water distribution coefficients have been measured at Genentech as part of an internship project. The data set is publicly available on Github, including the scripts used to analyze the results.