SEARCH
ARTICLE TYPES
Nonequilibrium candidate Monte Carlo is an efficient tool for equilibrium simulation
/Jerome P. Nilmeier, Gavin E. Crooks, David D. L. Minh, and John D. Chodera.
Proc. Natl. Acad. Sci. USA 108:E1009, 2011. [DOI] [PDF]
We present a significant generalization of Monte Carlo methods that provide an enormously useful tool for enhancing the efficiency of molecular simulations and enabling molecular design.
Keywords: NCMC; Monte Carlo; Metropolis-Hastings; acceptance rates; molecular dynamics
Dynamical reweighting: Improved estimates for dynamical properties from simulations at multiple temperatures
/John D. Chodera, William C. Swope, Frank Noé, Jan-Hendrik Prinz, Michael R. Shirts, and Vijay S. Pande.
J. Chem. Phys. 134:244107, 2011. [DOI] [PDF]
We describe how reweighing techniques can provide optimal estimates of temperature-dependent dynamical properties from simulations conducted at multiple temperatures.
Dynamical fingerprints: A theoretical framework for understanding biomolecular processes by combination of simulation and kinetic experiments
/Frank Noé, Sören Doose, Isabella Daidone, Marc Löllmann, Markus Sauer, John D. Chodera, and Jeremy C. Smith.
Proc. Natl. Acad. Sci. USA 108:4822, 2011. [DOI] [PDF]
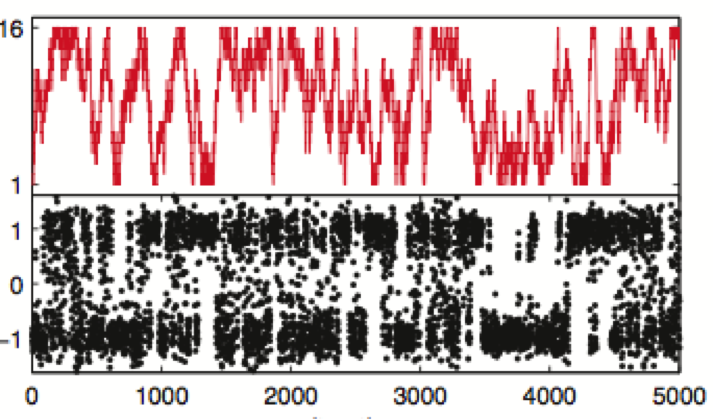
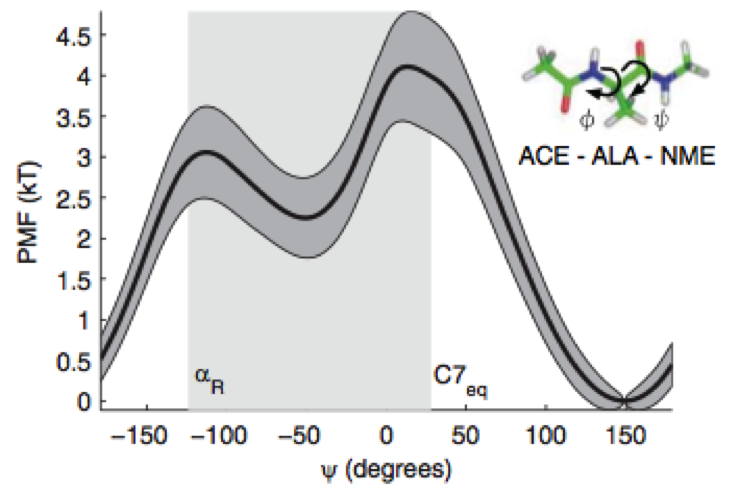
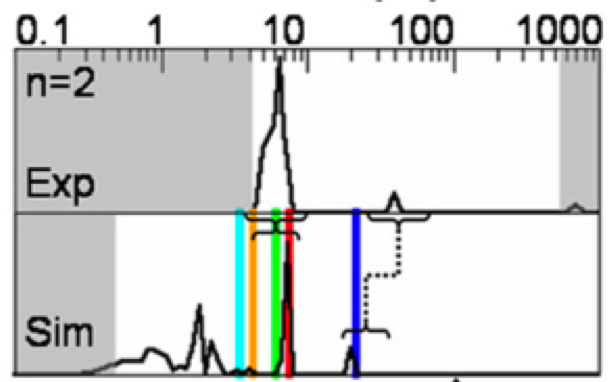
We present a new framework for comparing essential features of the dynamics between experiment and simulation to identify the kinetics processes contributing to individual relaxation timescales in perturbation-response or correlation spectroscopy experiments.
Estimating equilibrium ensemble averages using multiple time slices from driven nonequilibrium processes
/Estimating equilibrium ensemble averages using multiple time slices from driven nonequilibrium processes: Theory and application to free energies, moments, and thermodynamic length in single-molecule pulling experiments
David D. L. Minh and John D. Chodera
J. Chem. Phys. 134:024111, 2011. [DOI] [PDF]
We derive a new estimator for estimating equilibrium expectations from nonequilibrium experiments, and show how it can be used to estimate a variety of useful quantities in simulated single-molecule force spectroscopy experiments.
The mechanical properties of PCNA: Implications for the loading and function of a DNA sliding clamp
/Joshua L. Adelman, John D. Chodera, I. W. Kuo, Thomas F. Miller III, and Daniel Barsky.
Biophys. J. 98:3062, 2010. Featured on issue cover. [DOI] [PDF]
Molecular simulations of the PCNA clamp responsible for DNA polymerase processivity show a surprisingly small energetic penalty for the deformation required for clamp loading.
Current status of the AMOEBA polarizable force field
/Jay W. Ponder, Chuanjie Wu, Pengyu Ren, Vijay S. Pande, John D. Chodera, David L. Mobley, Michael J. Schnieders, Imran Haque, David S. Lambrecht, Robert A. DiStasio Jr., Martin Head-Gordon, Gary N. I. Clark, Margaret E. Johnson, and Teresa Head-Gordon.
J. Phys. Chem. B 114:2549, 2010. [DOI] [PDF]
A report on the status of the AMOEBA polarizable force field and its ability to reproduce a diverse set of physical chemical phenomenon to high accuracy.
Optimal estimators and asymptotic variances for nonequilibrium path-ensemble averages
/David D. D. L. Minh and John D. Chodera.
J. Chem. Phys. 131:134110, 2009. [DOI] [PDF]
We derive an optimal estimator and corresponding statistical uncertainties for inferring expectations of bidirectional nonequilibrium processes. These estimators have widespread applicability in single-molecule biophysical force-spectroscopy experiments and nonequilibrium molecular simulations.
Statistically optimal analysis of samples from multiple equilibrium states
/Michael R. Shirts and John D. Chodera.
J. Chem. Phys. 129:124105, 2008. [DOI] [PDF]
We present a highly general, statistically optimal approach for producing estimates of free energies and equilibrium expectations from multiple simulations that provably extracts all useful information from the data.
Keywords: Multistate Bennett acceptance ratio; MBAR; Bennett acceptance ratio; BAR; molecular dynamics; Monte Carlo; replica exchange
Predicting small-molecule solvation free energies: A blind challenge test for computational chemistry
/Anthony Nicholls, David L. Mobley, J. Peter Guthrie, John D. Chodera, and Vijay S. Pande.
J. Med. Chem. 51:769, 2008. [DOI] [PDF]
A blind evaluation of the accuracy of alchemical free energy methods for computing gas-to-water transfer free energies (solvation free energies) of small molecules demonstrates that modern forcefields are likely sufficiently accurate to be useful in drug design.
Accurate and efficient corrections for missing dispersion interactions in molecular simulations
/Michael R. Shirts, David L. Mobley, John D. Chodera, and Vijay S. Pande.
J. Phys. Chem. B 111:13052, 2007. [DOI] [PDF]
We identify a major source of systematic error in absolute alchemical free energy calculations of ligand binding and show how a simple procedure can inexpensively and accurately eliminate it.
Protein Folding by Zipping and Assembly
/S. Banu Ozkan, G. Albert Wu, John D. Chodera, and Ken A. Dill.
Proc. Natl. Acad. Sci. USA 104:11987, 2007. [DOI] [PDF]
A review of the utility of the proposed zipping and assembly mechanism for the concomitant formation of secondary and tertiary structure in protein folding for predicting folding pathways and native structures.
Automatic discovery of metastable states for the construction of Markov models of macromolecular conformational dynamics
/John D. Chodera*, Nina Singhal*, William C. Swope, Jed W. Pitera, Vijay S. Pande, and Ken A. Dill.
J. Chem. Phys. 126:155101, 2007. [DOI] [PDF]
Proposing one of the first automated algorithms for discovering kinetically metastable states of biomolecules from molecular simulations, this paper shows how many biomolecules can possess numerous distinct long-lived conformational states even though the the equilibrium populations of these states may too small for standard structural biology techniques to detect.
Long-time protein folding dynamics from short-time molecular dynamics simulations
/John D. Chodera, William C. Swope, Jed W. Pitera, and Ken A. Dill.
Multiscale Model. Simul. 5:1214, 2006. [DOI] [PDF]
We show how the long-time dynamics of biomolecular systems can be recapitulated from statistics collected from short molecular simulations sampling transitions between kinetically metastable states.